To learn more, please visit the YouTube Help Center: https://www.youtube.com/help
Get the latest international news and world events from around the world.
Ten Fourier and Laplace transformation
Here I solved ten Fourier and Laplace problem.laplace transformation.
inverse laplace transformation calculator.
laplace transformation calculator.
laplace transformation table.
inverse laplace transformation.
define laplace transformation.
laplace transformation chart.
laplace transformation differential equations.
laplace transformation examples.
laplace transformation calculator with steps.
how to do a laplace transformation.
application of laplace transformation.
laplace transformation application.
laplace transformation all formula.
laplace transformation ableitung.
laplace transform analysis gives.
laplace transform application in real life.
laplace transform all formulas pdf.
laplace transform and its applications pdf.
laplace transform and its properties.
laplace transform applications in engineering.
laplace transform and fourier series.
advantages of laplace transformation.
laplace transformation of e^at.
laplace transformation questions and answers.
laplace transformation of sin at.
laplace fourier and z transformation pdf.
laplace transformation book pdf.
laplace transform basic formula.
laplace transform basics.
laplace transform bessel function.
laplace transform by definition.
laplace transform btech notes.
laplace transform bsc 2nd year.
laplace transform bsc 2nd year pdf.
laplace transform by differentiation.
laplace transformation book.
laplace transformation bildfunktion.
transformation de laplace bibmath.
laplace transformation berechnen.
differentialgleichung laplace transformation beispiel.
differential equation by laplace transformation.
laplace transformation calculator wolfram.
laplace transformation cosh.
laplace transform circuit analysis.
laplace transform calculator piecewise.
laplace transform calculator with solutions.
laplace transform circuit analysis questions and answers.
laplace transform control systems.
laplace transformation of cosat.
inverse laplace transformation calculator with steps.
transformation de laplace exercices corrigés.
transformation de laplace exercices corrigés pdf.
transformation de laplace cours pdf.
laplace transformation definition.
laplace transformation derivative.
laplace transformation differentialgleichung.
laplace transform differential equation calculator.
laplace transform delta function.
laplace transform differential equations examples pdf.
laplace transform derivative formula.
laplace transform dirac delta.
define inverse laplace transformation.
diskrete laplace transformation.
doetsch handbuch der laplace-transformation.
dgl mit laplace transformation lösen.
derivatives of laplace transformation.
daniel jung laplace transformation.
differentialgleichung laplace transformation.
laplace transformation explained.
laplace transformation engineering mathematics.
laplace transformation elektrotechnik.
laplace transformation examples pdf.
laplace transformation engineering.
laplace transform examples and solutions pdf.
laplace transform examples and solutions.
laplace transform exercises.
laplace transform electrical circuit analysis.
eigenschaften der laplace transformation.
elektrotechnik laplace transformation.
einseitige laplace transformation.
e funktion laplace transformation.
eigenschaften laplace transformation.
laplace transformation formula.
laplace transformation formula sheet.
laplace transformation formula pdf.
laplace transformation for differential equation.
laplace transformation for dummies.
laplace transformation functions.
laplace transform formula list.
laplace transform first shifting theorem.
laplace transform final value theorem.
Transcending the Brain? AI, Radical Brain Enhancement and the Nature of Consciousness
Human Rights, Ethics, and Artificial Intelligence: Challenges for the next 70 Years of the Universal Declaration.
Susan schneider, university of connecticut, department of philosophy.
Transcending the Brain? AI, Radical Brain Enhancement and the Nature of Consciousness.
The views expressed in this video are those of the speaker(s) at the time of recording and do not necessarily reflect those of the Carr-Ryan Center for Human Rights or Harvard Kennedy School. These perspectives have been presented to encourage debate on important public policy challenges.
The Self as Software: Transcending and Enhancing the Brain
“The Self as Software: Transcending and Enhancing the Brain” presented by Susan Schneider. Co-sponsored by Cognitive Science, Computing Sciences, and Philosophy

Claude Fable 5 and Claude Mythos 5
While Mythos 5 remains largely unconstrained for restricted government and trusted enterprise partners, Fable 5 is wrapped in a sophisticated safety perimeter. If Fable 5 detects a prompt drifting toward high-risk vectors—like cyberwarfare exploits, advanced biology, or chemical synthesis—it doesn’t just give a generic “I can’t answer that” error. Instead, the query seamlessly falls back to Claude Opus 4.8 (Anthropic’s next-most capable model) to handle the response safely.
Today we’re launching Claude Fable 5: a Mythos-class1 model that we’ve made safe for general use.
Fable 5’s capabilities exceed those of any model we’ve ever made generally available. It is state-of-the-art on nearly all tested benchmarks of AI capability, showing exceptional performance in software engineering, knowledge work, vision, scientific research, and many other areas. The longer and more complex the task, the larger Fable 5’s lead over our other models.
Releasing a model this capable comes with risks. Without safeguards, Fable 5’s capabilities in areas like cybersecurity could be misused to cause serious damage. We’ve therefore launched the model with safeguards that mean queries on some topics will instead receive a response from our next-most-capable model, Claude Opus 4.8. To release the model both safely and quickly, we’ve tuned these safeguards conservatively—they’ll sometimes catch harmless requests, though they trigger, on average, in less than 5% of sessions. With more capable models arriving in the coming months, we’re working to improve our safeguards and reduce false positives as quickly as we can.

Researchers trigger sleep’s restorative effect in parts of the awake brain
Scientists from the University of Wisconsin-Madison have successfully replicated some of the restorative effects of deep sleep in awake mice by artificially inducing slow-wave brain activity. Using optogenetics to control specific neurons, researchers triggered localized cortical activity that mimics the NREM sleep phase responsible for synaptic homeostasis and the reorganization of neural connections. This targeted stimulation significantly reduced signs of fatigue and improved memory retention and cognitive performance in the mice following prolonged wakefulness. While the researchers caution that this technique is not a substitute for natural sleep, the findings suggest that localized neural stimulation can effectively preserve brain function during extended periods of wakefulness. Future research aims to explore whether similar cognitive benefits can be achieved in humans through non-invasive methods, such as transcranial electrical stimulation.
NIH-funded study in animals offers new details about how the brain resets during sleep.
By inducing specific patterns of activity in small portions of the brain in awake mice, researchers supported by the National Institutes of Health (NIH) have triggered a recalibration of neural connections that normally only occurs during sleep. This new approach offset the effects of sleep deprivation in memory tasks and revealed features of sleep that are key to its restorative effect.
“What we’re essentially doing is forcing sleep in a local region of the brain. While that part is solidifying memories and restoring learning capacity, other parts stay aware/vigilant and connected to environment,” said corresponding author Chiara Cirelli, M.D., Ph.D., a professor of psychiatry at the University of Wisconsin-Madison. “Dolphins do something similar, sleeping with only one brain hemisphere at a time.”
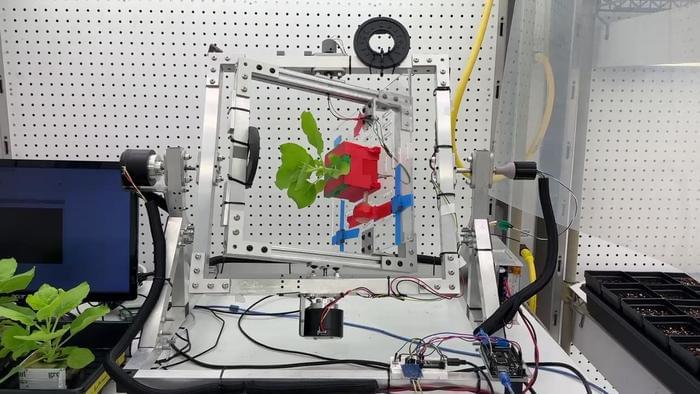

Space Plants Could Be Future Pharmacies for Astronauts
Scientists have successfully tested a non-destructive method to harvest life-saving medicines from plants under simulated space conditions, enabling on-demand drug production for long-duration missions. [ https://www.labroots.com/trending/space/30644/space-plants-f…tronauts-2](https://www.labroots.com/trending/space/30644/space-plants-f…tronauts-2)
How can plants help produce pharmaceuticals for future astronauts? This is what a recent study published in npj Science of Plants hopes to address as a team of scientists from the University of California San Diego (UCSD) investigated using plants to produce drugs for astronauts to treat a variety of ailments. This study has the potential to help scientists, mission planners, and astronauts develop new methods for addressing medical concerns on long-term space missions.
For the study, the researchers examined how cowpea mosaic virus (CPMV) could be produced under space-like conditions, including a vacuum environment, microgravity, using a centrifuge, the latter of which is commonly used in space for science experiments. CPMV is a plant virus-based compound that has been found to treat cancer while also possessing immunotherapy characteristics. The primary motivation behind the study was to address how to provide medical treatments to astronauts on long-term space missions without relying on Earth supplies. In the end, the researchers found that CPMV could successfully be extracted without harming the plants.
The study notes, “The combination of process-level and host-level optimization facilitates sustainable CPMV production under the constrained conditions of long-duration space missions while also offering practical advantages for terrestrial biomanufacturing.”

New pilot plant converts unsorted plastic waste into oil in 30 mins
A mobile pilot plant has been designed to convert various types of plastic waste into oil.
Developed by the Catalysis Engineering Group at the University of Amsterdam (UvA), the Solvothermal Liquefaction (STL) process uses a potent mix of solvent, heat, catalysts, and intense pressure to cook mixed plastic waste back into oil.
Interestingly, the resulting dark brown oil contains the precise molecules needed to remake brand-new, virgin plastic, thereby closing the recycling loop.