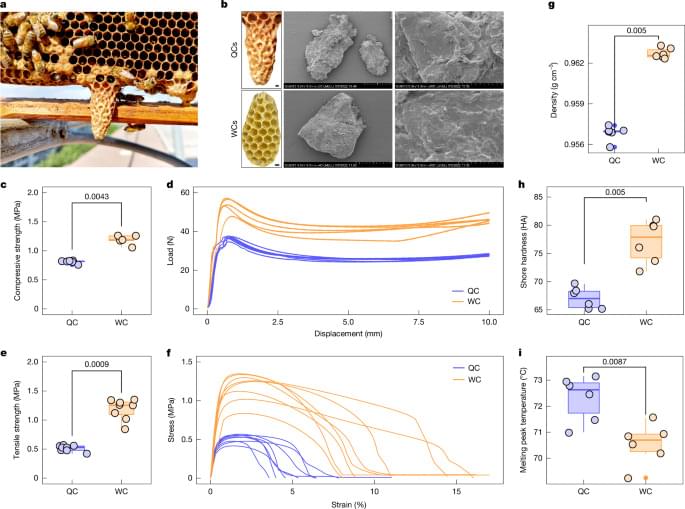
Nature — Worker bee construction behaviour actively engineers a physicochemical niche that is crucial for queen development in honey bees.

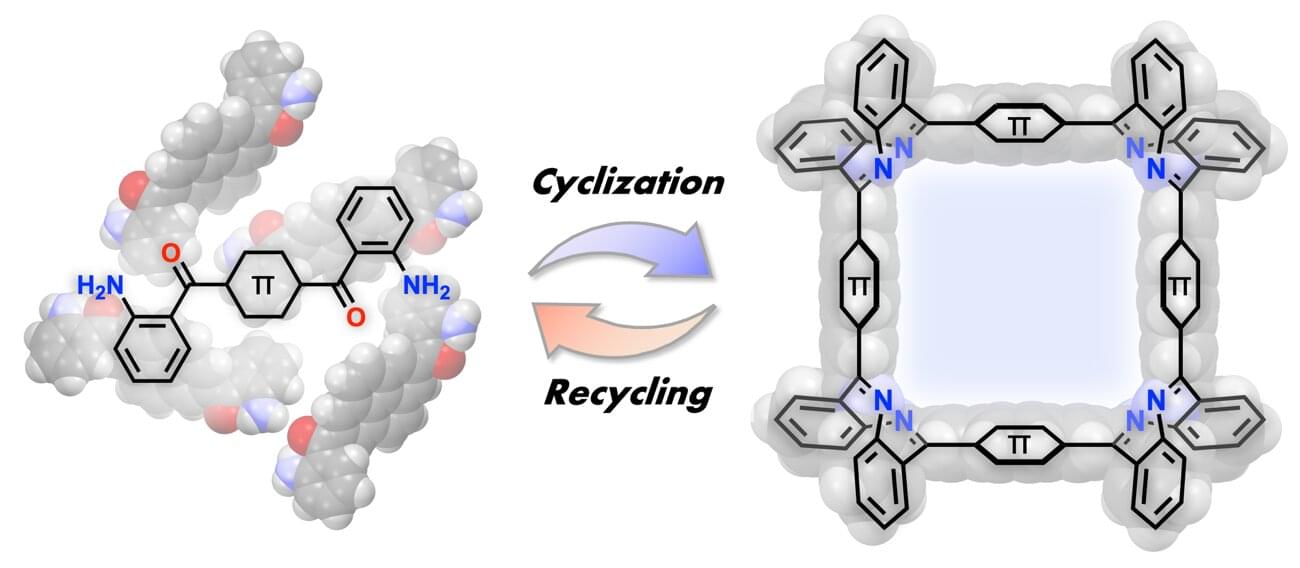
A research group has developed a new method for selectively synthesizing three-dimensional macrocycles,⁽¹⁾ in which four panels are arranged in a square, by connecting planar π-conjugated molecules⁽²⁾ at right angles.
This method is applicable to a wide variety of π-conjugated molecules and allows the size of the internal cavity to be designed. Furthermore, the resulting square macrocycles exhibit acid responsiveness, reversibly changing color under the action of a mild acid, while acid-mediated hydrolysis enables the starting monomers to be recovered in high yield—realizing a sustainable molecular synthesis that reverts to and regenerates the starting materials. The originality of this work lies in having a single imine bond play three roles: creating the shape, responding to stimuli and reverting back.
These research results were published in the Journal of the American Chemical Society on Monday, June 1, 2026. The team includes Associate Professor Yasutomo Segawa and Assistant Professor Takashi Harimoto at the Institute for Molecular Science (National Institutes of Natural Sciences) and the Graduate University for Advanced Studies (SOKENDAI).
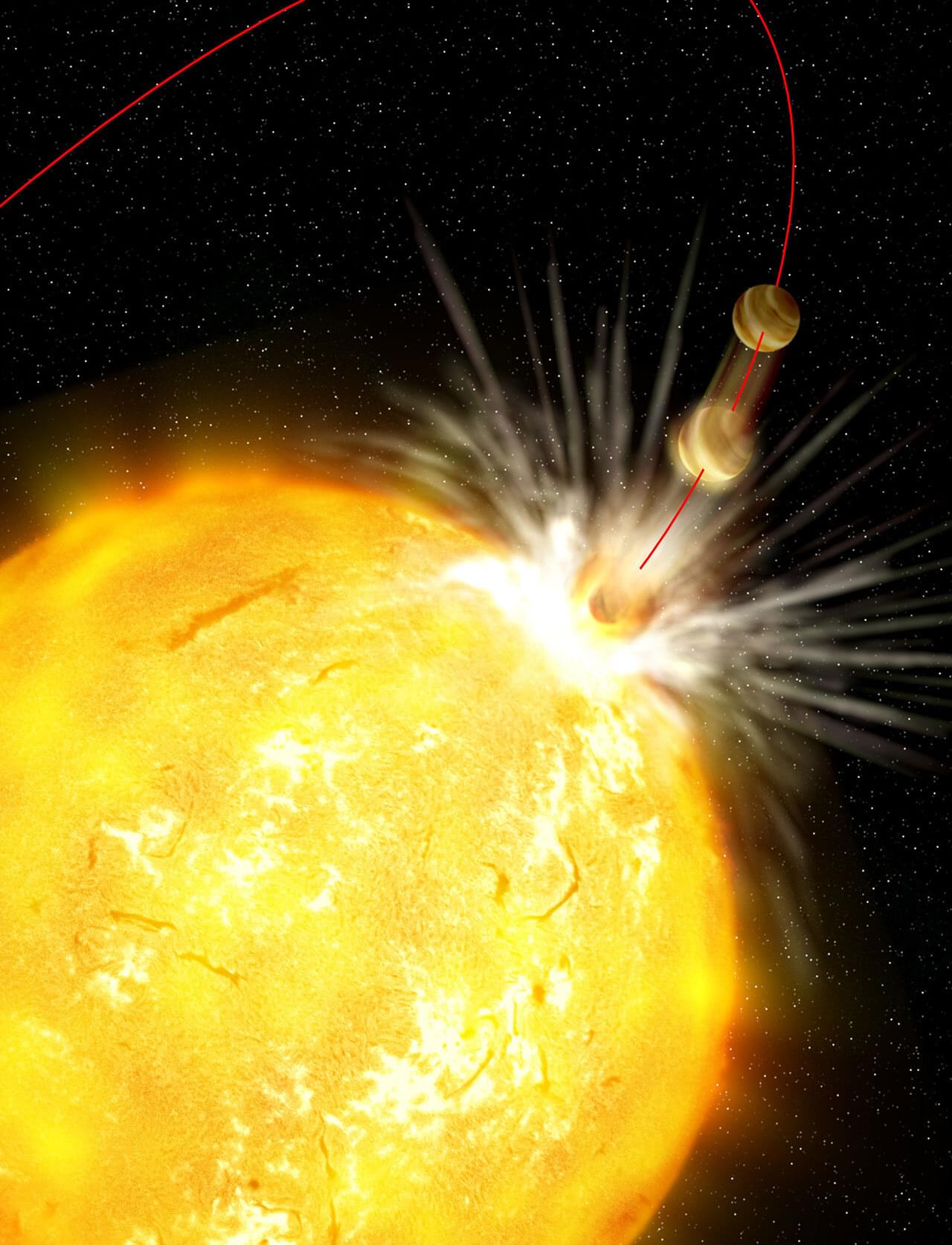

Astronomers have investigated a puzzling binary star system in which two stars that may have formed together now show dramatically different chemical compositions. The new study, uploaded to the arXiv preprint server on May 29, hints at the possibility that one of the stars may have swallowed its own planets.
Generally, in binary systems, the two stars form from the same molecular cloud and, as a result, have the same age and chemical composition. Any differences in their metallicity, astronomers say, hint at an event involving mass transfer or engulfment of planetary components or other internal processes. HD 81,809 is one such peculiar system in which the stars are both sun-like G stars but are at different stages of evolution.
The primary star, HD 81809A, has crossed the main-sequence phase, depleted its hydrogen fuel in the core but hasn’t turned into a giant star yet—it is now a subgiant. On the other hand, the secondary star, HD 81809B, is still a main-sequence star. It has lithium enrichment and there is a difference in iron content between the two stars—the primary is metal-poor with an iron abundance of −0.57 dex, while the secondary has roughly solar metallicity around 0.00 dex.

If everything that happens in the world ultimately comes down to the behavior of fundamental particles, it would seem that other entities, from cells to human beings, from currencies to financial markets, aren’t really causing anything at all—that they are just shadows cast by patterns at the most fundamental level. But philosopher David Yates argues this conclusion is wrong. The whole affects the parts, and higher-level structures don’t just describe what is happening at lower levels in more convenient terms—they actively shape what is possible. This means that chemists, biologists, psychologists, and economists aren’t chasing shadows. They are studying structures that genuinely shape how the world unfolds.
In 1974, Jerry Fodor published a seminal paper titled ‘Special Sciences’, in which he argued for an intuitive and compelling picture of the relationship between fundamental physics and higher-level sciences such as biology, psychology and economics. Our world, according to Fodor, is arranged hierarchically, with fundamental physical particles at the bottom, combining to form molecules, which combine to form cells, which combine to form complex organisms, some of which have mental states, among them humans, who combine to form complex societies. The sciences are likewise arranged, with physics at the bottom, followed by chemistry, biology, physiology, neuroscience, psychology, sociology and economics. Now it is vanishingly unlikely, says Fodor, that things that share e.g. psychological or economic properties, also share some property specifiable in the language of physics or other lower-level sciences.
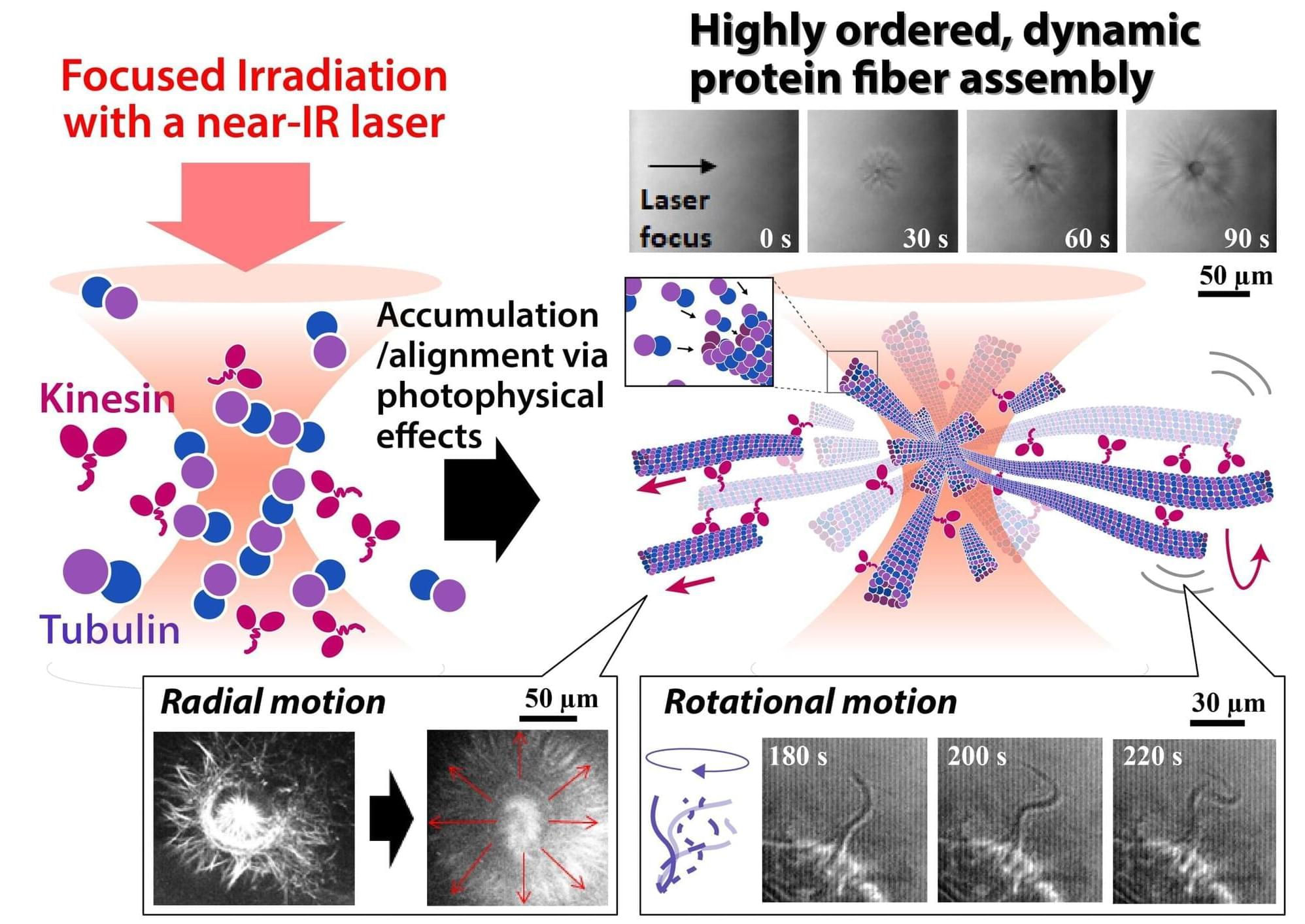
Networks of protein fibers play important roles in living cells. To understand the dynamical behavior of these networks, model networks are needed to perform in vitro studies. However, fabrication of protein networks similar to those in cells has proved difficult, as current methods could affect the biological function of these proteins—ultimately impacting our understanding of any findings.
Now, researchers at The University of Osaka and Saitama University have used a laser beam to precisely fabricate a network of protein fibers. Their discovery was recently reported in Advanced Science.
The shape of living cells is determined by an internal network of protein fibers called a cytoskeleton. The cytoskeletal structure is dynamic, as the key nodes for cell function shift over time. One such cell function can be witnessed with motor proteins, which convert chemical energy into mechanical work. These proteins walk along cytoskeletal tracks to drive muscle contraction and transport components across the cell.
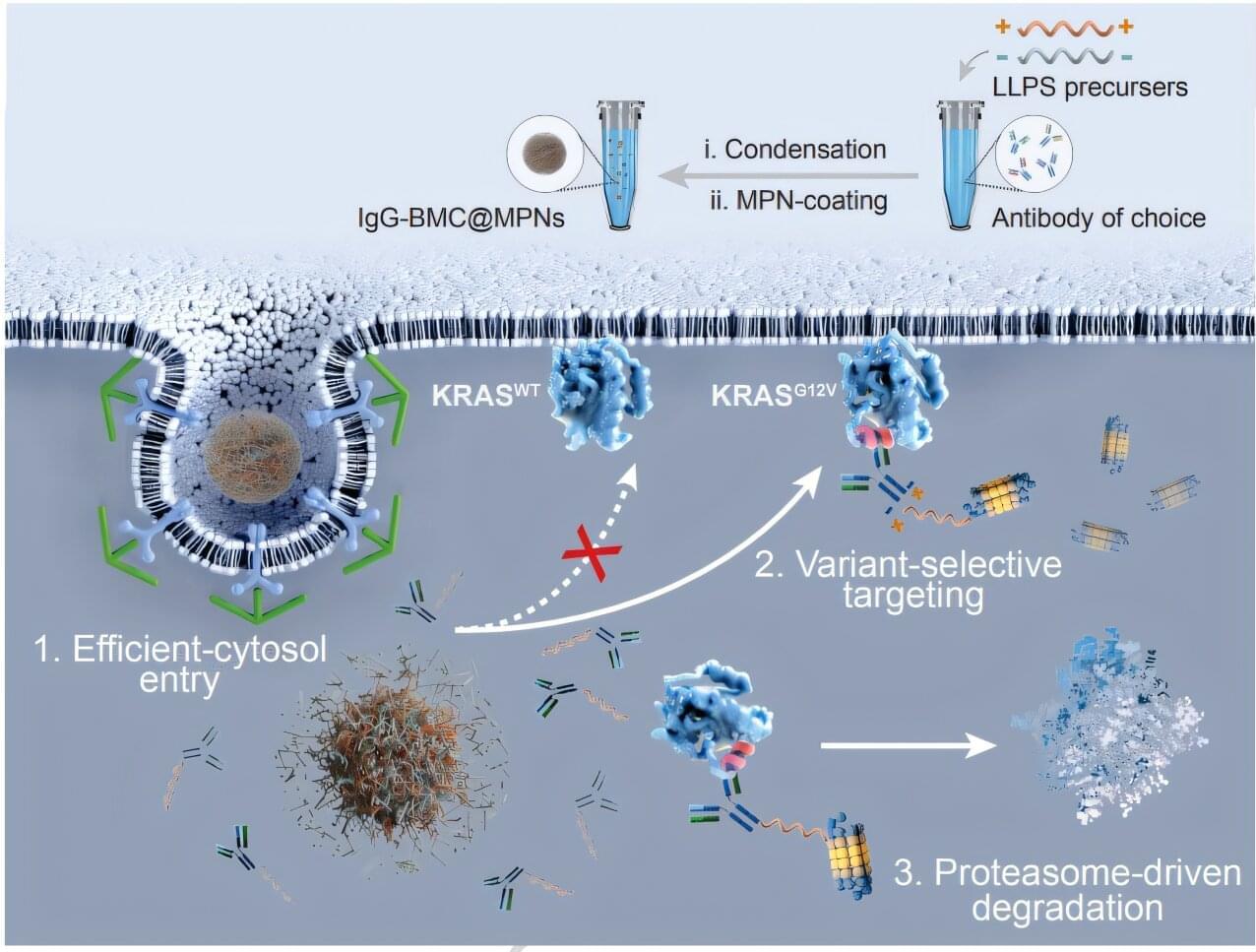
Northwestern Medicine scientists have developed a novel synthetic biomolecular condensate that can degrade intracellular disease-causing proteins, providing a framework for new therapeutic approaches for a wide range of diseases, as detailed in a recent study published in Nature Communications.
Shana Kelley, Ph.D., the Neena B. Schwartz Professor of Chemistry, Biomedical Engineering, and Biochemistry and Molecular Genetics and the president of the Chan Zuckerberg Biohub Chicago, was senior author of the study.
Targeted protein degradation is an emerging therapeutic strategy that harnesses cells’ own degradation machinery to clear disease-causing proteins. However, achieving this degradation process across different cell types has remained a challenge due to subtle variations in protein structure.
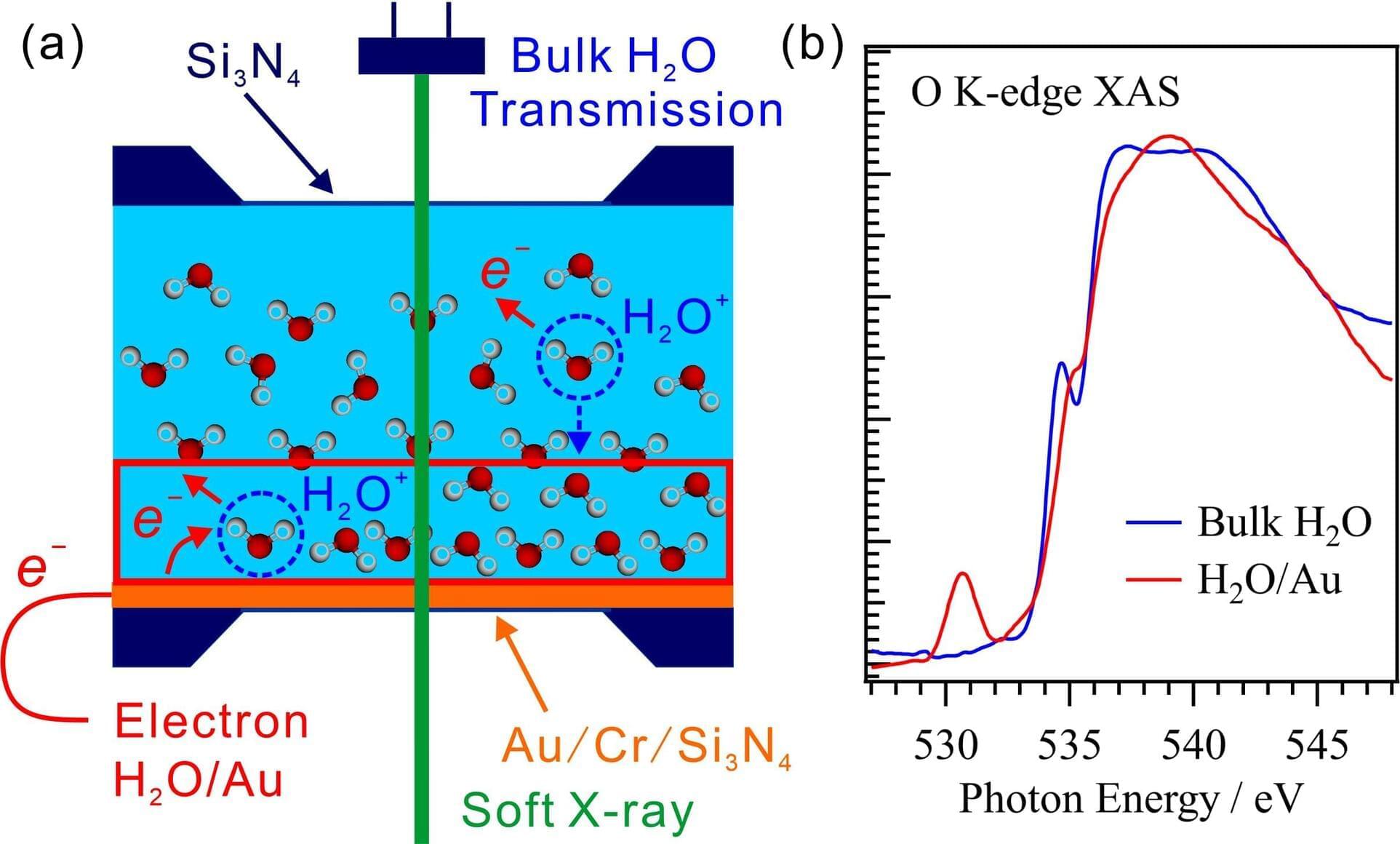
Researchers have developed a method for making simultaneous soft X-ray absorption spectroscopy (XAS) measurements of solid-liquid interfaces and bulk liquids. By controlling the thickness of the liquid layer, they obtained the O K-edge XAS spectrum of bulk H2O from a liquid H2O layer on a thin Au film using the transmission method, and they used the electron-yield method to obtain the XAS spectrum of the H2O/Au interface by measuring the drain currents from the Au surface following soft X-ray absorption. This method for obtaining simultaneous XAS measurements of solid-liquid interfaces and bulk liquids can be utilized to investigate the mechanisms of a variety of catalytic, electrochemical, and biological reactions involving solid-liquid interfaces.
Water molecules at solid-liquid interfaces play important roles in various catalytic, electrochemical, and biological reactions. Soft X-ray absorption spectroscopy (XAS) is an element-specific method for investigating the electronic structures of liquid water and organic molecules. In this study, the researchers developed a method for simultaneously obtaining XAS measurements of a solid-liquid interface, using the electron-yield method, and of the bulk liquid, using the transmission method. The paper is published in the Journal of Synchrotron Radiation.
In the present work, they measured the XAS spectra while precisely controlling the thickness of the liquid layer in the range from 20 nm to 40 μm in a liquid cell for the transmission of soft X-rays. The XAS spectra acquired in transmission mode are derived mainly from the bulk liquid because the contributions from the solid-liquid interfaces are smaller than those from the bulk liquid. In contrast, the XAS spectra of solid-liquid interfaces are obtained by detecting Auger electrons, which originate mostly from those interfaces because they escape only from shallow depths.
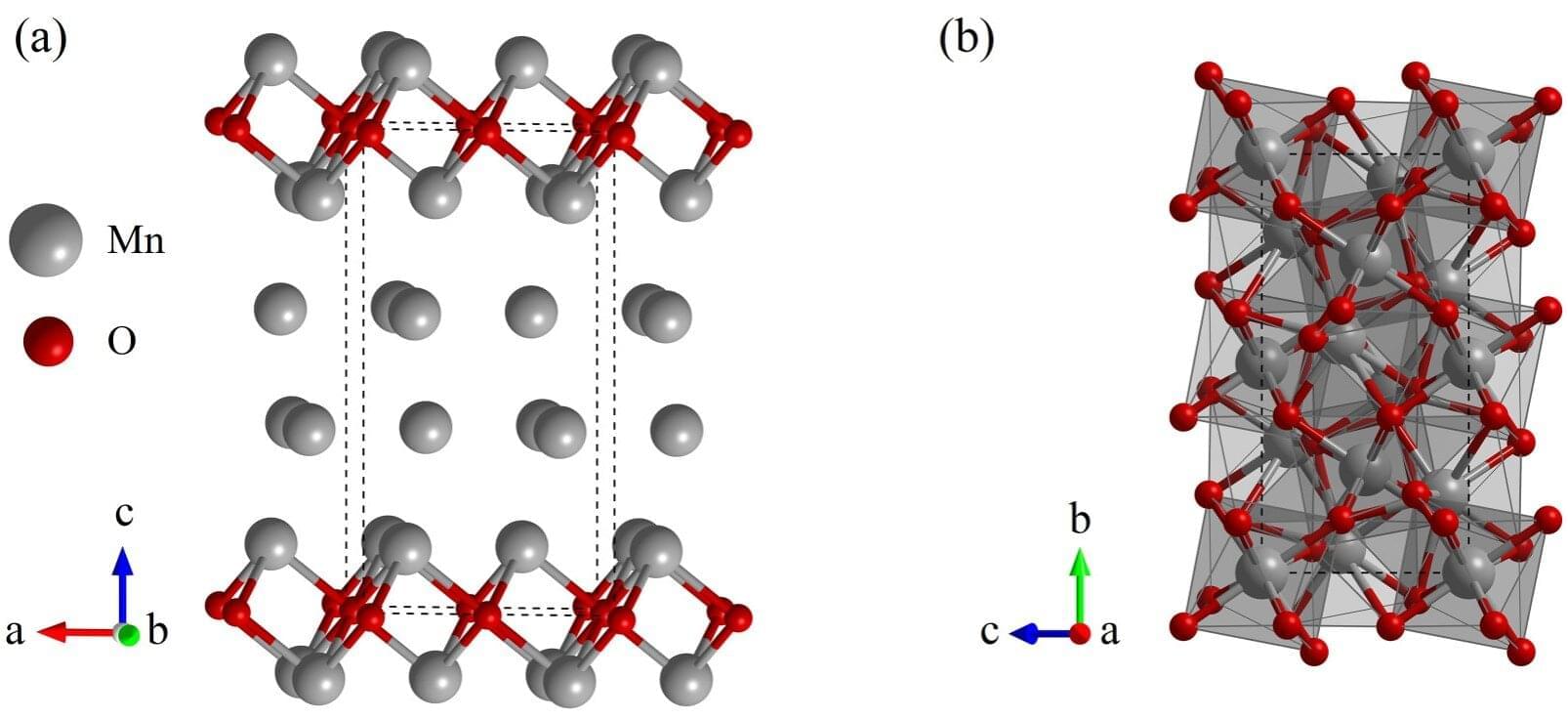
Scientists know that manganese, in its various oxide forms, plays a significant role in Earth’s geochemical cycles. However, the exact forms of manganese, their abundance and the mechanisms behind these cycles that occur in Earth’s deep, high-pressure interior are not well understood. But, a recent study, published in Physical Review B, reports on a newly discovered manganese rich compound that might help shed light on manganese’s behavior in Earth’s interior and explain why seismic waves slow down in certain regions.
While Earth’s mantle mostly consists of oxygen, magnesium, silicon, and iron, other elements, like manganese, also play an important role. Manganese oxides, such as MnO, Mn3O4, Mn2O3, MnO2, are known to exist in Earth’s interior and have been studied in the context of their stability in the high-pressure conditions of Earth’s mantle, but researchers think there may be additional manganese oxides involved.
These compounds have the ability to react with other compounds (and oxidize) depending on the surrounding pressure and temperature. They often act as powerful, pressure-sensitive redox agents, actively participating in deep-Earth geochemical cycling by reacting with and oxidizing subducted iron-bearing minerals.
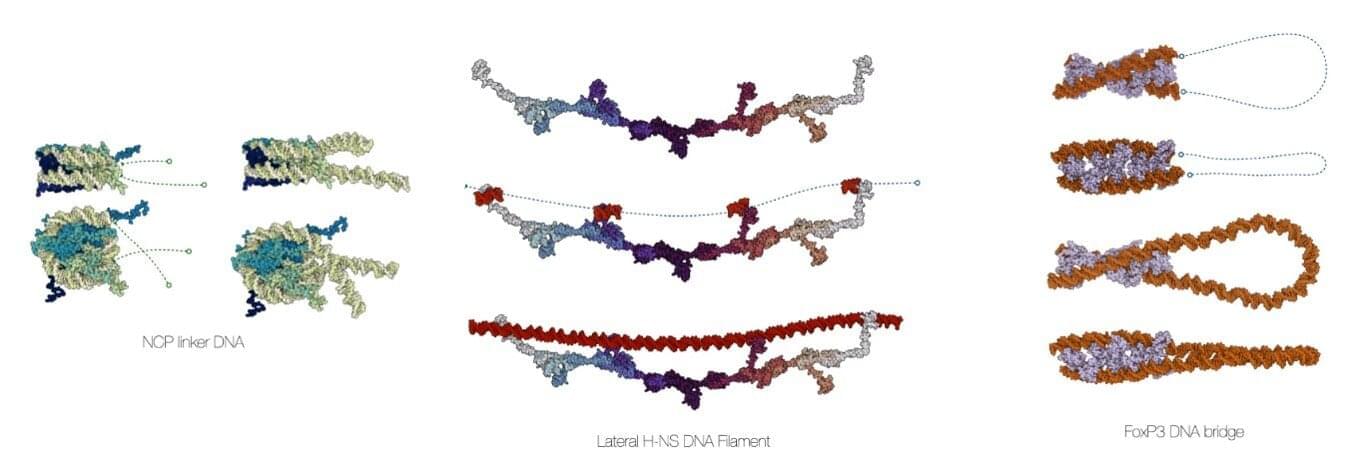
Computational chemists at the University of Amsterdam’s Van ‘t Hoff Institute for Molecular Sciences have developed a comprehensive software suite to create accurate models of DNA in biomolecular assemblies. Called MDNA, the user-friendly molecular modeling toolkit helps biochemists, molecular biologists, bioinformaticians, and biophysicists to visualize and analyze DNA structures and perform accurate simulations.
The development of the MDNA suite, led by associate professor Jocelyne Vreede, has been presented in a paper in Nucleic Acids Research.
The software is open-source and publicly available through Figshare and Github. It is easily accessible, providing inspiration to any scientist with an interest in DNA. It has been thoroughly tested by students in mathematics, chemistry and biology, some of whom had hardly any programming experience.
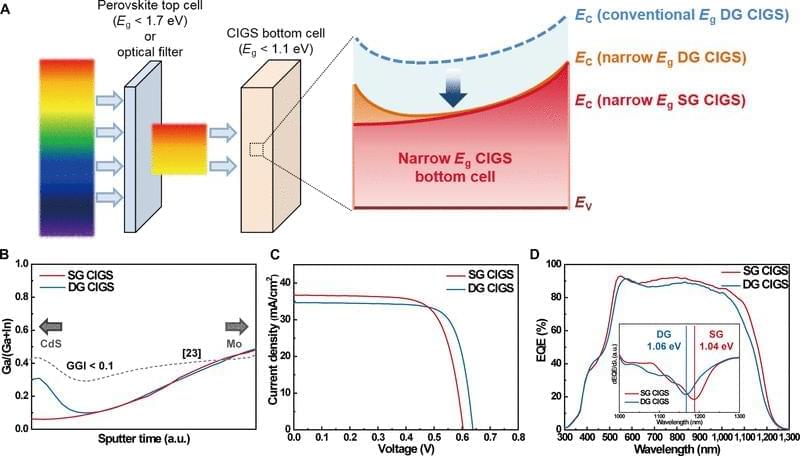
This study examined the potential of narrow-bandgap (Perovskite-based tandem solar cells are a promising photovoltaic (PV) technology to exceed the Shockley–Queisser limit of single-junction solar cells. Perovskite/Si tandem solar cells have been intensively studied, demonstrating a record power conversion efficiency (PCE) of 34.6% [1]. In contrast, the certified record PCE of perovskite/Cu(In, Ga)Se2 (CIGS) tandem solar cells remains 24.6% with a reported efficiency of 24.9% [1, 2]. Theoretical calculations for double-junction tandem solar cells using a detailed balance model indicate that the bandgap (Eg) combinations of 1.12 eV (for a bottom cell) and 1.70 eV (for a top cell) or 0.90 to 1.04 eV (for a bottom cell) and 1.58 to 1.67 eV (for a top cell) can yield a maximum theoretical tandem efficiency [3, 4]. Wide-bandgap perovskite (with Eg equal to or greater than 1.7 eV) has been actively studied for tandem application with Si (Eg = 1.12 eV), the most successful solar cell technology to date as a bottom cell. However, previous studies have shown that wide-bandgap perovskite suffers from substantial open-circuit voltage (VOC) loss due to halide segregation [5], and the maximum PCEs of single-junction perovskite cells have been produced by perovskite with Eg between 1.52 and 1.63 eV [6– 8]. The bandgap of CIGS can be tuned between 1.01 and 1.68 eV by adjusting the Ga/(Ga+In) (GGI) ratio and through tuning of bandgap grading profile [9]. Employing a narrow-bandgap CIGS close to 1.00 eV as a bottom cell is advantageous to use the most efficient, conventional bandgap perovskite as the top cell. Therefore, unlike Si, the bandgap tunability of CIGS offers an opportunity for perovskite/CIGS to attain a greater ultimate performance than perovskite/Si tandem solar cells. Han et al. [10] introduced a thick indium-doped tin oxide (ITO) recombination layer to bury the intrinsic surface roughness of CIGS, followed by chemical mechanical polishing to prepare a smooth surface for the subsequent solution process of perovskite, attaining a certified PCE of 22.4%. Albrecht and coworkers have improved the PCE of perovskite/CIGS tandem solar cells by modifying the hole transport layer (HTL). In their earlier work, a NiOx/PTAA bilayer was utilized to form a uniform HTL on CIGS bottom cells. Recently, a self-assembled monolayer such as 2PACz and Me-4PACz was used, which can enhance the device performance of single-junction perovskite solar cell and its perovskite/CIGS tandem counterpart, achieving a certified PCE of 24.2% [2, 11 – 13].
Most recent studies on perovskite/CIGS tandem solar cells have focused on optimizing the perovskite top cell. In contrast, all CIGS bottom cells include an absorber with a double-graded (DG) bandgap profile optimized around the bandgap of ~1.1 eV. The DG bandgap profile has been adapted primarily for CIGS absorbers prepared by thermal evaporation, which has resulted in high-performing CIGS solar cells with PCEs up to 23.4% [14], and it has proven to be an effective strategy for enhancing performance, optimized for “single-junction” CIGS; however, it has not been determined whether DG would be the ideal configuration for tandem applications. Kim et al. [15] used single-graded (SG) CIGS with a bandgap close to 1.0 eV, where the band grading is only formed on the backside of the absorber. They employed dual alkali post-deposition treatment (PDT) with KF and CsF, demonstrating a CIGS solar cell with a PCE of 20.