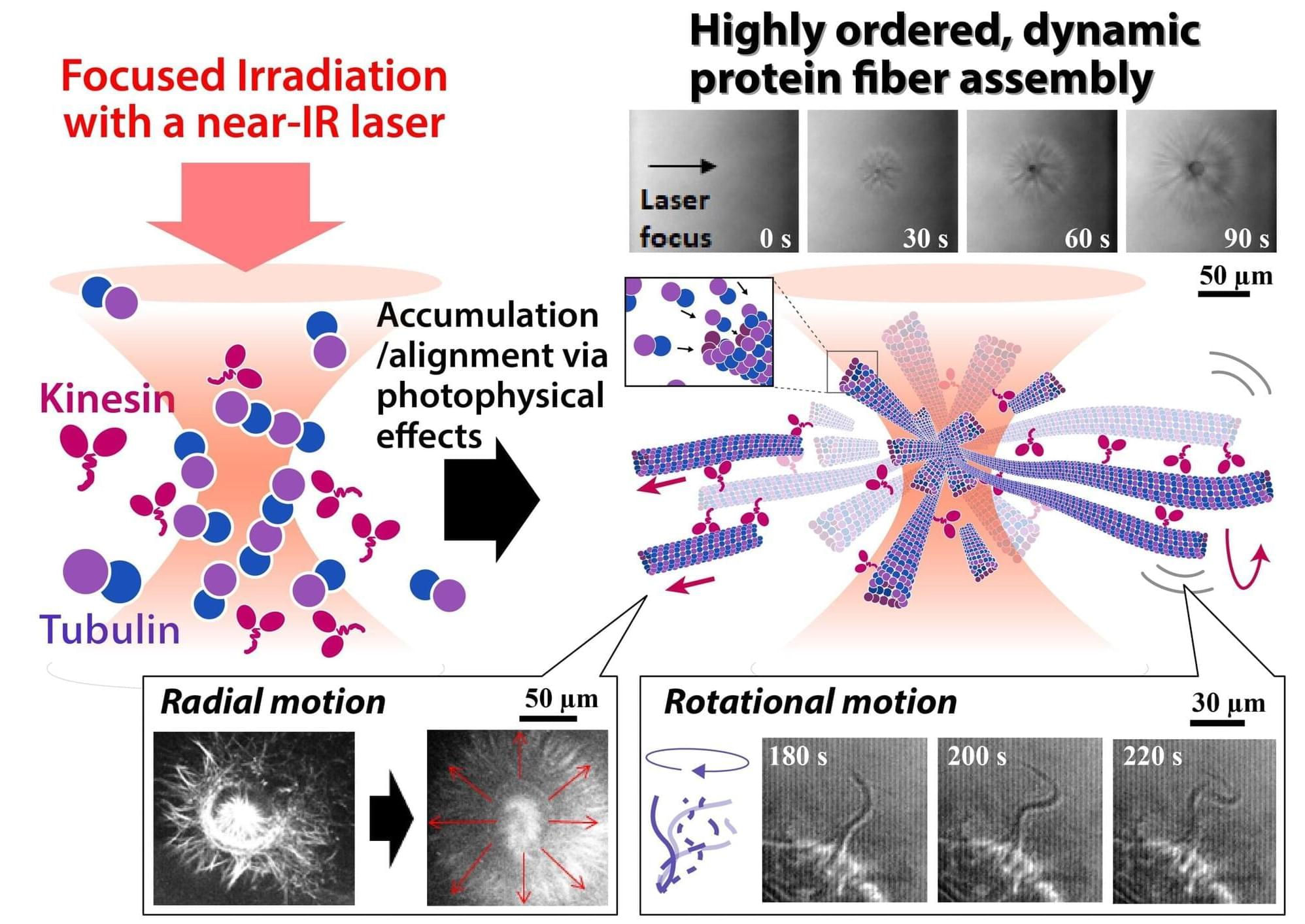
Networks of protein fibers play important roles in living cells. To understand the dynamical behavior of these networks, model networks are needed to perform in vitro studies. However, fabrication of protein networks similar to those in cells has proved difficult, as current methods could affect the biological function of these proteins—ultimately impacting our understanding of any findings.
Now, researchers at The University of Osaka and Saitama University have used a laser beam to precisely fabricate a network of protein fibers. Their discovery was recently reported in Advanced Science.
The shape of living cells is determined by an internal network of protein fibers called a cytoskeleton. The cytoskeletal structure is dynamic, as the key nodes for cell function shift over time. One such cell function can be witnessed with motor proteins, which convert chemical energy into mechanical work. These proteins walk along cytoskeletal tracks to drive muscle contraction and transport components across the cell.