Learn more about the holographic principle and why some theoretical physicists think the universe could be a hologram.
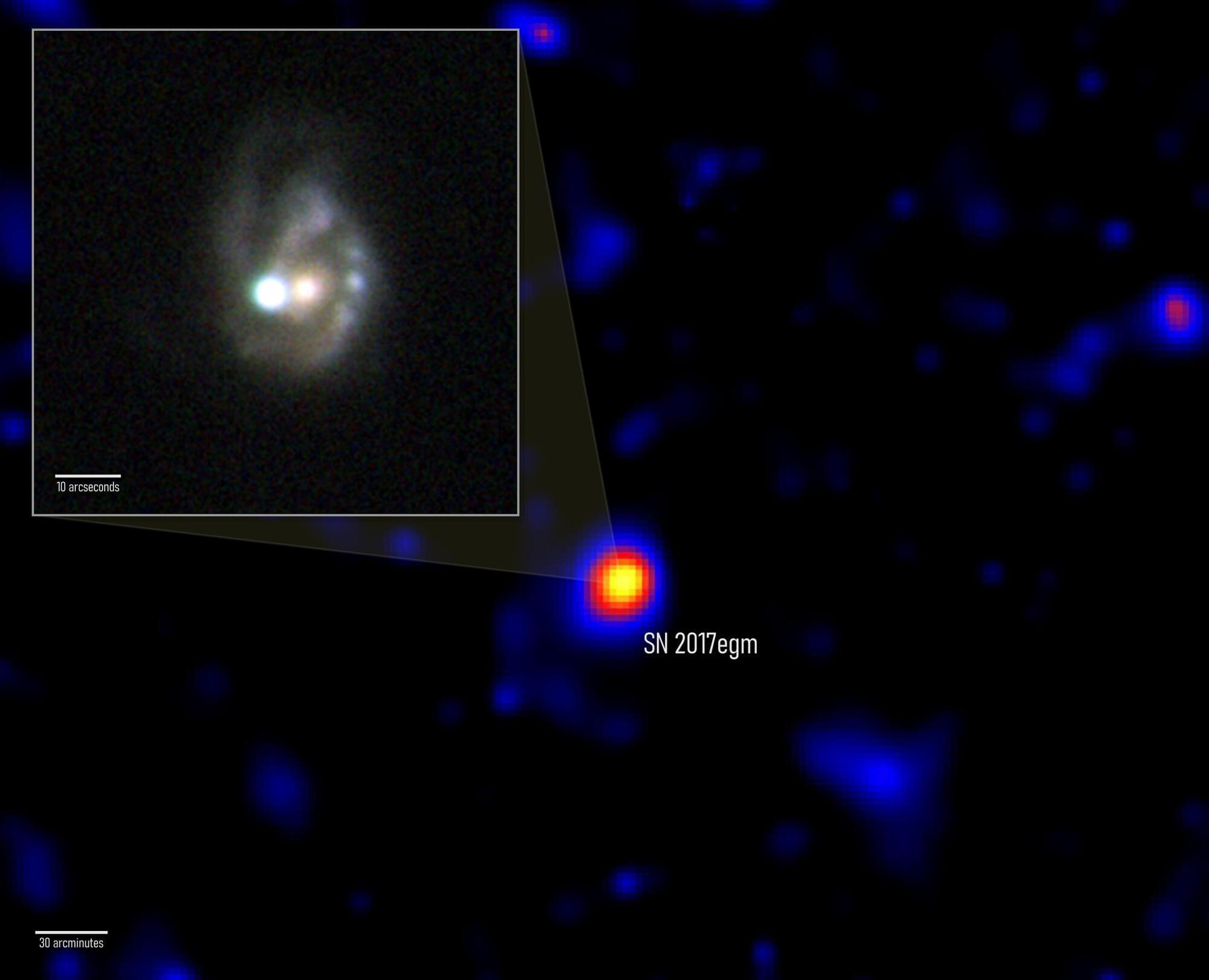



NASA’s Fermi telescope has detected what may be the first confirmed gamma-ray signal from a superluminous supernova — one of the most extreme explosions in the universe. Scientists believe the blast was powered by a rapidly spinning magnetar, an exotic neutron star with unbelievably strong magnetic fields. The event, called SN 2017egm, erupted 440 million light-years away and may help explain why some supernovae become extraordinarily bright.
NASA’s Fermi Gamma-ray Space Telescope may have finally uncovered what powers some of the brightest stellar explosions ever observed. After studying years of data, an international research team found strong evidence that a rare superluminous supernova was energized by an extremely magnetic neutron star formed during the star’s collapse.
The Fermi mission is part of NASA’s network of observatories designed to track changing events across the universe and help scientists better understand how cosmic phenomena work.
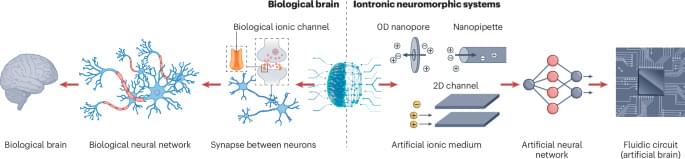
In the brain, memory involves release of neurotransmitters and transport of ions through nanoconfined channels. This Perspective discusses how nanofluidic memristors emulate this confined ion transport, highlighting the materials, design strategies and challenges involved in developing brain-inspired computing technologies.

face_with_colon_three I still think that ceramics would be very useful to stop the need for global mining operations that rely heavily on rare materials when they can make the same chip from ceramics.
To be shown at ECTC 2026, May 26–29 in Orlando, USA, the new substrate technology delivers superior rigidity and circuit miniaturization for next-gen data centers, AI, and ASIC packaging.
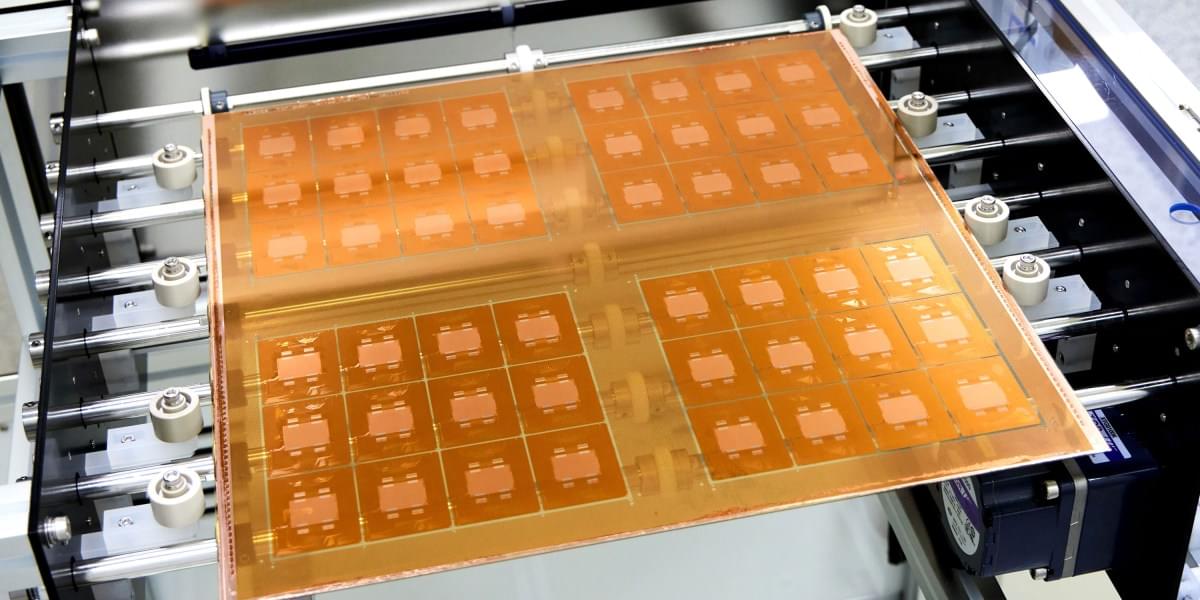

The idea is to use glass as the substrate, or layer, on which multiple silicon chips are connected. This form of “packaging” is an increasingly popular way to build computing hardware, because it lets engineers combine specialized chips designed for specific functions into a single system. But it presents challenges, including the fact that hardworking chips can run so hot they physically warp the substrate they’re built on. This can lead to misaligned components and may reduce how efficiently the chips can be cooled, leading to damage or premature failure.
“As AI workloads surge and package sizes expand, the industry is confronting very real mechanical constraints that impact the trajectory of high-performance computing,” says Deepak Kulkarni, a senior fellow at the chip design company Advanced Micro Devices (AMD). “One of the most fundamental is warpage.”
That’s where glass comes in. It can handle the added heat better than existing substrates, and it will let engineers keep shrinking chip packages—which will make them faster and more energy efficient. It “unlocks the ability to keep scaling package footprints without hitting a mechanical wall,” says Kulkarni.
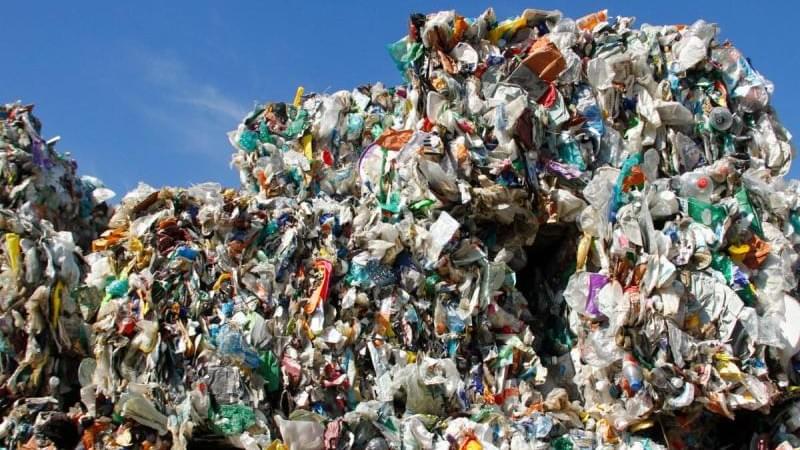
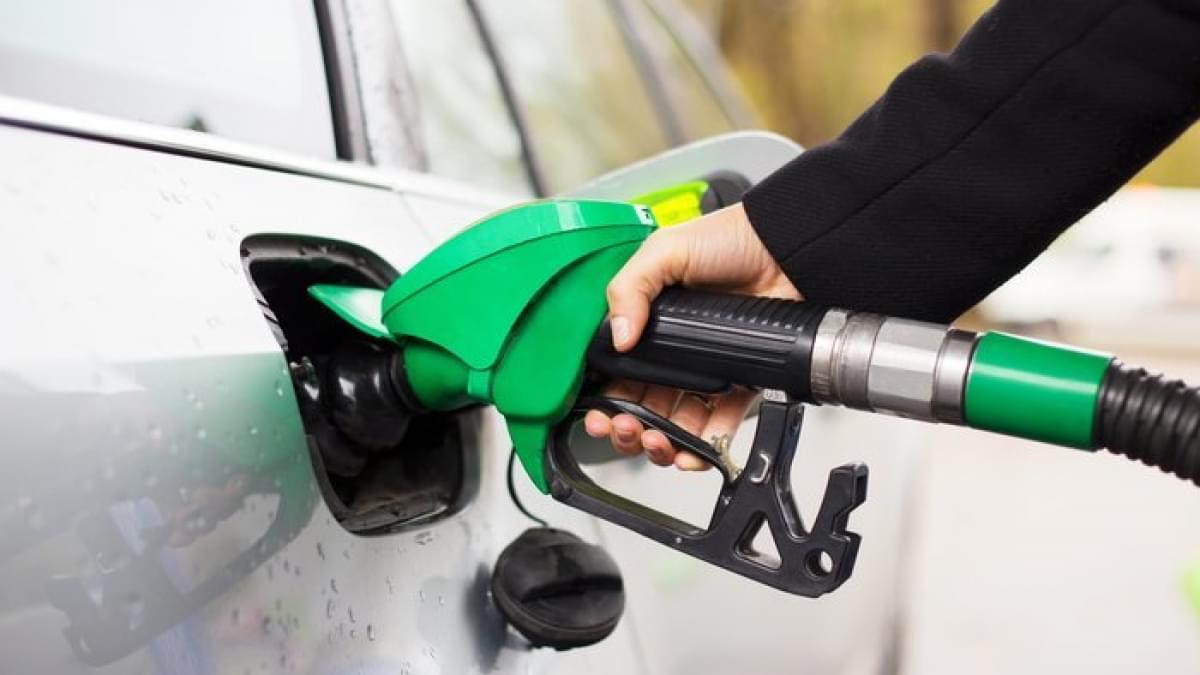



KU Leuven, Belgium bioscience engineers have developed a roadmap, so to speak, for industrial cellulose gasoline.
The bioscience engineers already knew how to make gasoline in the laboratory from plant waste such as sawdust. In 2014, at KU Leuven’s Centre for Surface Chemistry and Catalysis, the researchers succeeded in converting sawdust into building blocks for gasoline.
A chemical process made it possible to convert the cellulose – the main component of plant fibers – in the sawdust into hydrocarbon chains. These hydrocarbons can be used as an additive in gasoline. The resulting cellulose gasoline is a second-generation biofuel.


Experimental attosecond science is built around the ability to generate and control light flashes lasting billionths of a billionth of a second. Such extreme pulses can be created through high harmonic generation (HHG), where an intense laser field drives electrons out of atoms or solids and then forces them back, releasing bursts of extreme ultraviolet radiation. Techniques like this have transformed our ability to observe electron motion on its natural timescale.
To extract information from such ultrafast processes, physicists often rely on attosecond interferometry. By combining a strong laser field with a weaker second colour, different electron trajectories are made to interfere, imprinting timing and phase information onto the emitted harmonics. Over recent years, these schemes have become standard tools for attosecond metrology and spectroscopy.