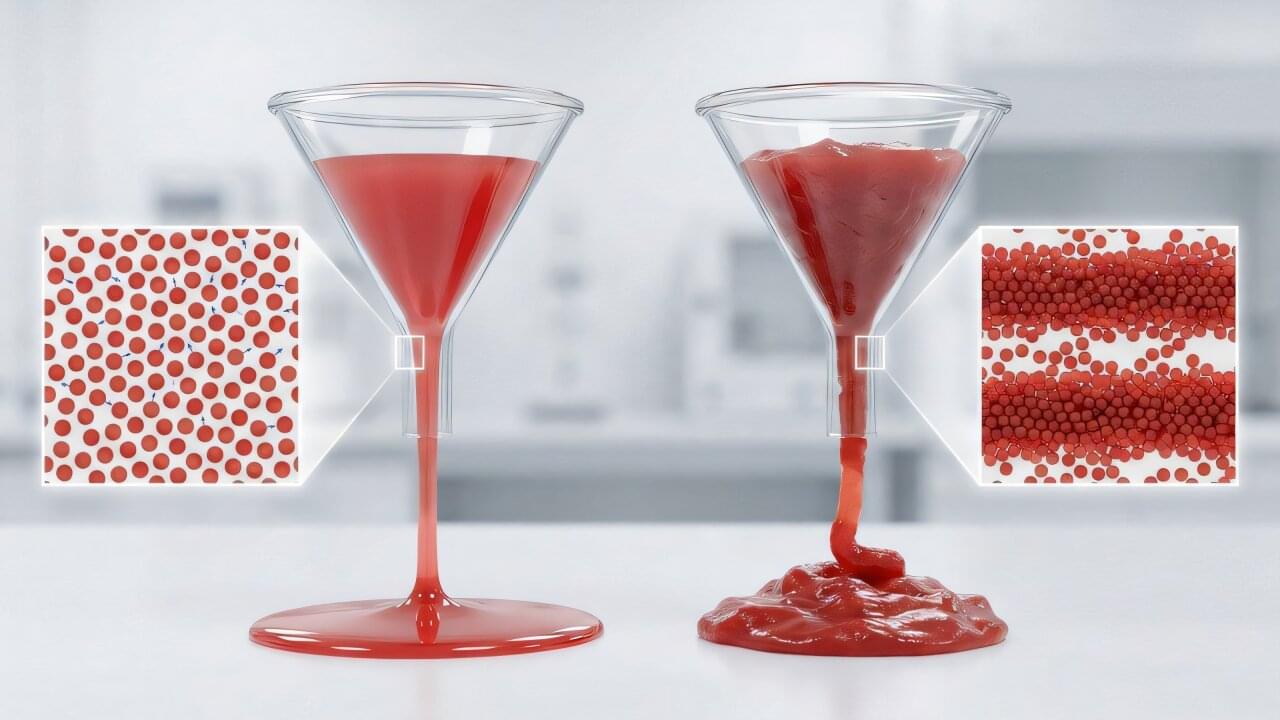
Sitting in a restaurant, you reach for the ketchup bottle, eyeing the basket of fries in front of you. You give the bottle a shake, then a tap. For a moment, nothing happens—the ketchup clings stubbornly to the glass. Then, all at once, it lets go and rushes out, sometimes in a steady stream, sometimes in a messy surge that threatens to flood the basket.
That awkward moment when ketchup stops behaving like a solid and suddenly starts flowing like a liquid is called “yielding.” Scientists see the same kind of behavior in many everyday and advanced materials, from toothpaste, paints and concrete to 3D-printing inks and electrodes used in next-generation batteries. Yet, what actually causes a material to hold its shape one moment and suddenly let go the next has been surprisingly hard to pin down, especially deep inside dense, opaque fluids where particle motion is difficult to see.