A study in mice and nematodes has investigated a potential new therapeutic approach that could help people with the genetic variant that predisposes them to Alzheimer’s disease.



Biopharmaceutical impurities such as host cell proteins can delay biologics development and production. These elements can have immunogenic effects in patients, forcing scientists to restart process development. Thus, detecting and removing biopharmaceutical impurities is necessary for maintaining drug efficacy and ensuring patient safety.
In this webinar brought to you by Cytiva, Andrew Hamilton and Joe Hirano will discuss how to identify, detect, and measure impurities in biologics manufacturing.
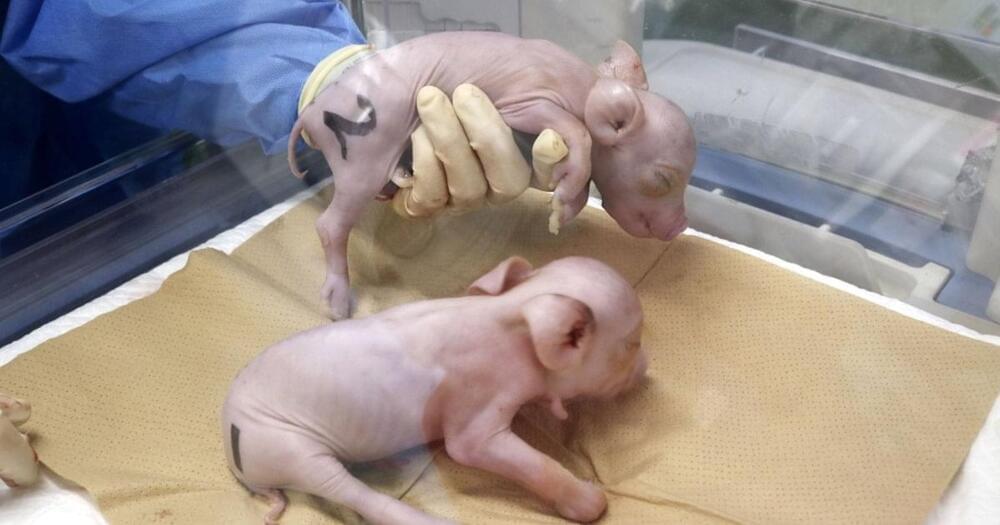
Japanese startup PorMedTec says it’s have cloned three piglets with the express purpose of having their organs be viable for transplantation to humans, without being rejected by the immune system.
The company imported gene-edited cells from a US biotech startup called eGenesis and used them to create genetically modified embryos, the Japan Times reports, which were then implanted into the uterus of a pig.
“The realization of xenotransplantation has been long awaited in Japan for several years, but it remained in the basic research stage because pigs that could withstand clinical application were still under development,” the company said in a statement.

Sounds like the video game/movie “The Last of Us” though there was a somewhat similar X Files episode as well before that. Though I doubt it’ll be a zombie plague, it could be like another pandemic someday or an issue such as a deadly fungal outbreak they had in Portland, Oregon before if I recall.
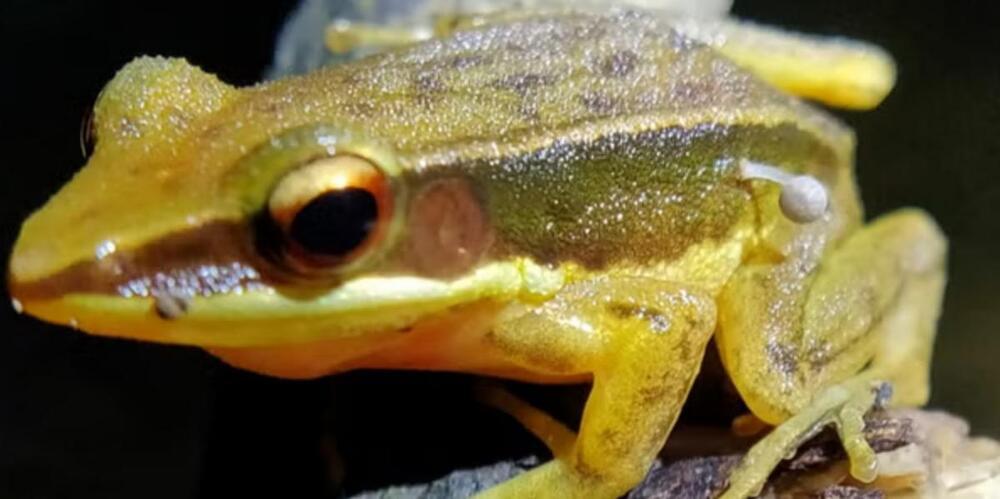
Scientists have been left concerned after making the surprise discovery of a frog with a small mushroom sprouting from its leg.
The amphibian was discovered in the foothills of India’s Western Ghats and researchers stated that it’s the first time a mushroom has been found growing on live animal tissue.
Researchers affiliated with the World Wildlife Fund released findings on the species, known as Rao’s intermediate golden-backed frog (Hylarana intermedia), in a new study published in the journal Reptiles and Amphibians.

Scentists have uncovered a fascinating link between ancient viruses and the development of myelination, the biological process crucial for the advanced functioning of the nervous system in vertebrates, including humans.
Scientists discovered a gene, ‘RetroMyelin,’ from ancient viruses, essential for myelination in vertebrates, suggesting viral sequences in early vertebrate genomes were pivotal for developing complex brains. This breakthrough in Cell unravels how myelination evolved, highlighting its significance in vertebrate diversity.
“We’ve used the mutations that give cancer cells their staying power to engineer what we call a ‘Judo T-cell therapy’ that can survive and thrive in the harsh conditions that tumors create,” said co-author Kole Roybal.
CAR-T cells: CAR-T cell therapy starts with doctors extracting T cells — a type of white blood cell — from a cancer patient’s blood. They then engineer the T cells to display proteins called “chimeric antigen receptors” (CARs) that bind to cancer cells.
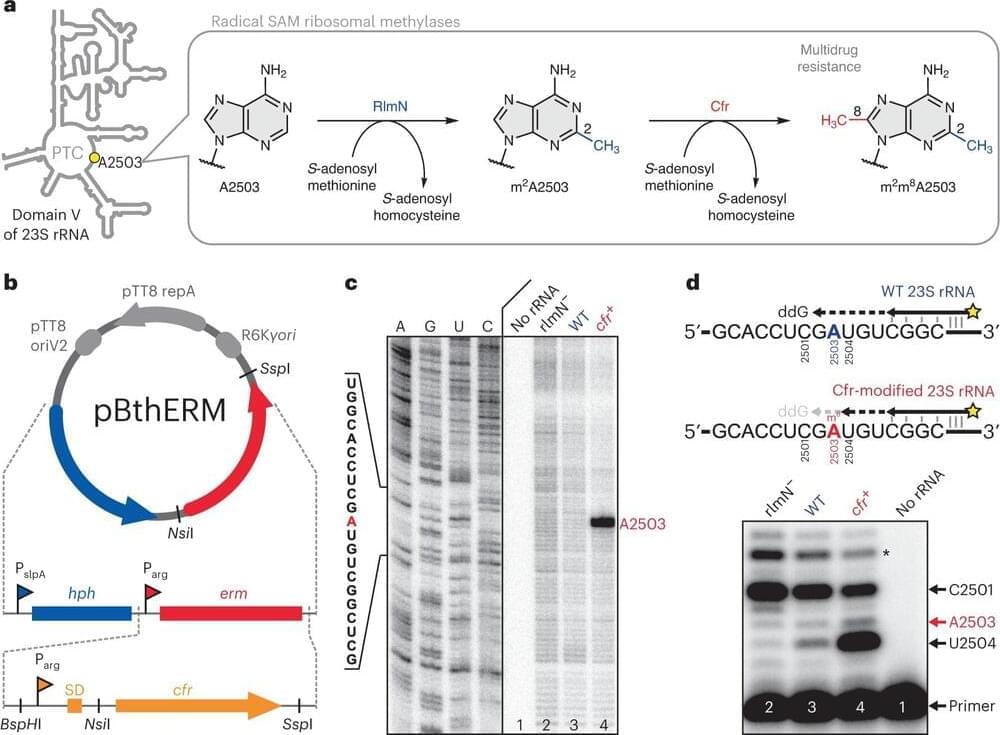
Scientists at the University of Illinois Chicago and Harvard University have developed an antibiotic that could give medicine a new weapon to fight drug-resistant bacteria and the diseases they cause.
The antibiotic, cresomycin, described in Science, effectively suppresses pathogenic bacteria that have become resistant to many commonly prescribed antimicrobial drugs.
The promising novel antibiotic is the latest finding for a longtime research partnership between the group of Yury Polikanov, associate professor of biological sciences at UIC, and colleagues at Harvard. The UIC scientists provide critical insights into cellular mechanisms and structure that help the researchers at Harvard design and synthesize new drugs.
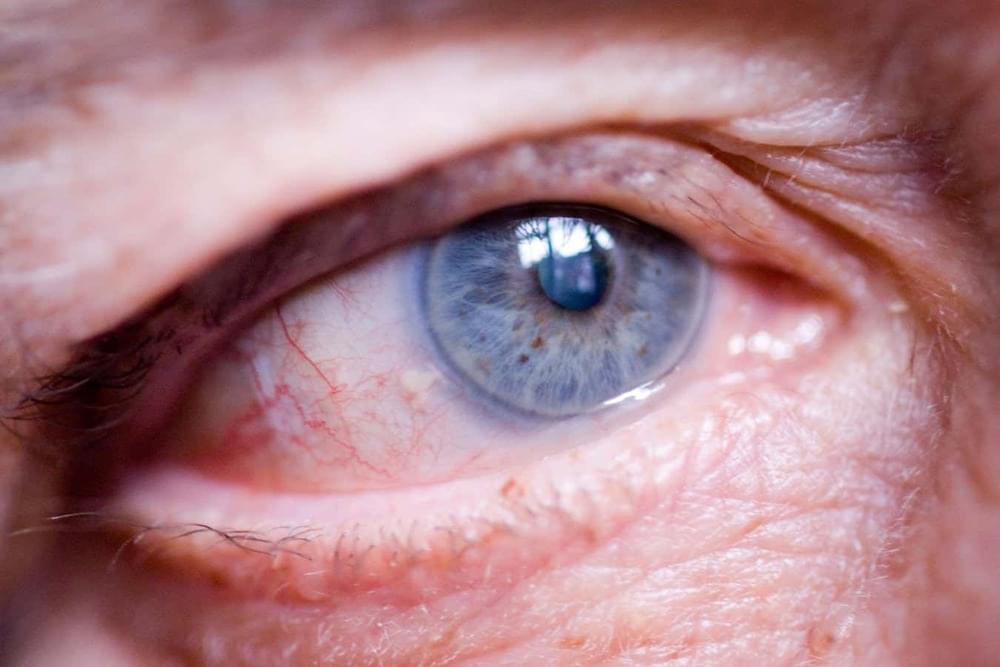
The optic nerve has been partly regenerated in mice, raising hopes for treating blindness caused by conditions such as glaucoma.
This video explores 20 emerging technologies and their future. Watch this next video about the 10 stages of AI: • The 10 Stages of Artificial Intelligence.
🎁 5 Free ChatGPT Prompts To Become a Superhuman: https://bit.ly/3Oka9FM
🤖 AI for Business Leaders (Udacity Program): https://bit.ly/3Qjxkmu.
☕ My Patreon: / futurebusinesstech.
➡️ Official Discord Server: / discord.
💡 Future Business Tech explores the future of technology and the world.
Examples of topics I cover include:
• Artificial Intelligence & Robotics.
• Virtual and Augmented Reality.
• Brain-Computer Interfaces.
• Transhumanism.
• Genetic Engineering.
SUBSCRIBE: https://bit.ly/3geLDGO
Disclaimer:
Some links in this description are affiliate links.
As an Amazon Associate, I earn from qualifying purchases.
This video explores 20 emerging technologies and their future. Other related terms: ai, artificial intelligence, future business tech, future technology, future tech, future business technologies, future technologies, artificial general intelligence, artificial superintelligence, superintelligence, future city, radical life extension, crisp, quantum computer, neuralink, humanoid robot, generative ai, starlink, nanotechnology, smart cities, mixed reality, autonomous vehicles, blockchain, lab grown meat, smart home, fusion power, space tourism, artificial wombs, etc.
Scientists at The University of Texas at Austin have discovered how an aggressive and deadly form of leukemia fuels its growth. In an experimental study, they were able to curb the cancer’s growth without harming healthy cells. The finding provides clues for future drug developers about how to increase the effectiveness of one type of chemotherapy.
The study, led by Xiaolu Cambronne of the Department of Molecular Biosciences, in collaboration also with researchers at Dell Medical School and in the Department of Nutritional Sciences, is published in Cell Metabolism.
Acute myeloid leukemia (AML) is an aggressive cancer that starts in the blood-forming cells of the bone marrow. Known for rapid expansion, the cancer kills approximately 11,000 Americans each year. Most of the cases of AML occur in adults over 65, a population that often responds poorly to aggressive treatments, such as bone marrow transplants, and thus has limited options.